After completion of reaction (TLC: 8:1/hexanes:EtOAc), the solution was washed with water (1 × 100 ml), 1 M aqueous HCl (3 × 100 ml), 0.5% aqueous NaHCO 3 (3 × 100 ml) and brine (1 × 100 ml). The reaction mixture was then heated to reflux for 4 h. NBS (18.5 g, 105 mmol) was added to a stirred solution of 4-methyl acetophone (13.4 g, 100 mmol) in 400 ml of carbon tetrachloride, followed by the addition of 2′,2′-azobisiosbutyronitrile (AIBN 0.43 g, 2.5 mmol).

N-bromosuccinimide (NBS) was recrystallized before usage. Synthesis of p-Acetyl-(±)- l-Phenylalanine ( 13). Importantly, this tRNA-synthetase pair is orthogonal to its counterparts for the common 20 amino acids i.e., the orthogonal synthetase (and only this synthetase) aminoacylates the orthogonal tRNA (and only this tRNA) with the unnatural amino acid only, and the resulting acylated tRNA inserts the unnatural amino acid only in response to the amber codon.
coli in response to (and only in response to) an amber non-sense codon. coli in the form of p-acetyl- l-phenylalanine, a tRNA-synthetase pair was evolved that is capable of inserting this amino acid site-specifically into proteins in E. To genetically encode this functional group in E. Although present in cofactors ( 10), metabolites ( 11), and as a posttranslational modification to proteins ( 12), this important functional group is absent from the side chains of the common amino acids. Moreover, the unique reactivity of the keto group allows it to be selectively modified with hydrazide and hydroxylamine derivatives in the presence of the other amino acid side chains ( 7– 9). The keto group is ubiquitous in organic chemistry, and participates in a large number of reactions from addition reactions to aldol condensations. coli, and that the unique reactivity of the keto group can be used to modify proteins selectively in vitro with a wide variety of agents.
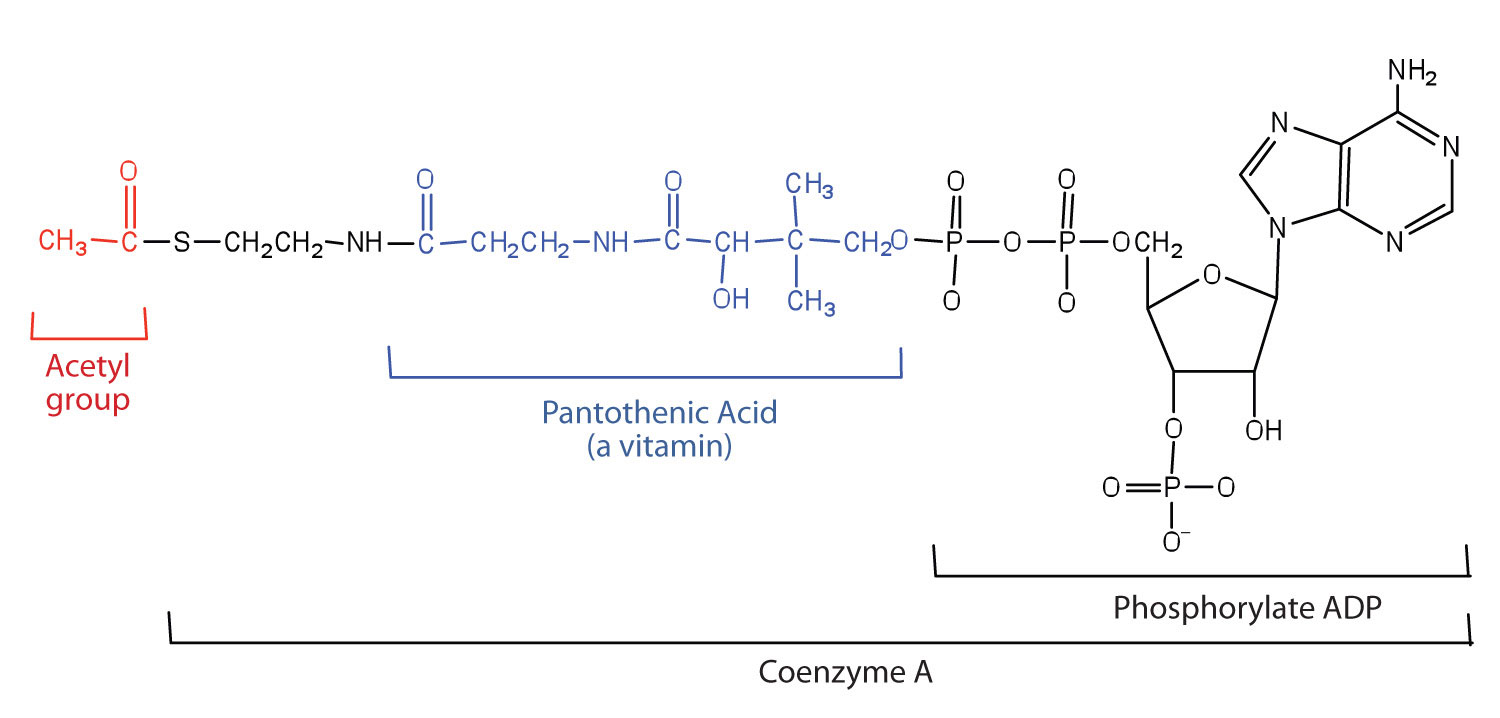
Acetyl functional group code#
We now demonstrate that this approach can be extended to add a keto-containing amino acid to the genetic code of E. Recently, we reported that, by adding new components to the translational machinery of Escherichia coli, one could site-specifically incorporate with high fidelity a number of unnatural amino acids ( 4– 6) into proteins in vivo. The ability to augment the genetically encoded amino acids with new amino acids, for example, amino acids with metal chelating, fluorescent, redox active, photoactive, or spin-labeled side chains, would significantly enhance our ability to manipulate the structures and functions of proteins and perhaps living organisms themselves.
The side chains of the common amino acids comprise a surprisingly limited number of functional groups: nitrogen bases, carboxylic acids and amides, alcohols, and a thiol group, the remainder being simple alkanes or hydrophobic groups. Only in rare cases are selenocysteine ( 1) or pyrrolysine ( 2, 3) added. The genetic codes of most known organisms encode the same common 20 amino acids as building blocks for the biosynthesis of proteins.
